12 Jan METAGENOMICS FOR DETECTION OF GREEN MOLD DISEASE DURING CHAMPIGNON CULTIVATION
Dr. Ljiljana Šašić Zorić, senior researcher, BioSense Institute
Andrijana Andrić, senior researcher, BioSense Institute
Mushrooms have been present in human nutrition due to their rich proteins, minerals and vitamins content. The most common cultivated species is the white button mushroom or champignon (Agaricus bisporus). The champignon is cultivated using specially prepared, partially composted organic substrate which functions as a growing medium and a nutrient source. The mycelium grows through compost by degrading the organic material to release necessary nutrients. Additionally, the champignon requires a layer of peat called “casing soil”, placed on the surface of the composted substrate to induce the development of the mushroom fruiting bodies and maintain compost environmental conditions. The compost and casing microbial communities are known to play important roles in the mushroom production process: (1) being an important source of nitrogen for mushroom mycelium, (2) releasing sugar residues through the degradation of the wheat straw in the composted substrate, (3) playing a critical role in inducing development of fruiting bodies and (4) acting as pathogens by parasitising the mushroom mycelium. Thus, study of microbial community diversity is of great importance for understanding and improvement of champignon production.
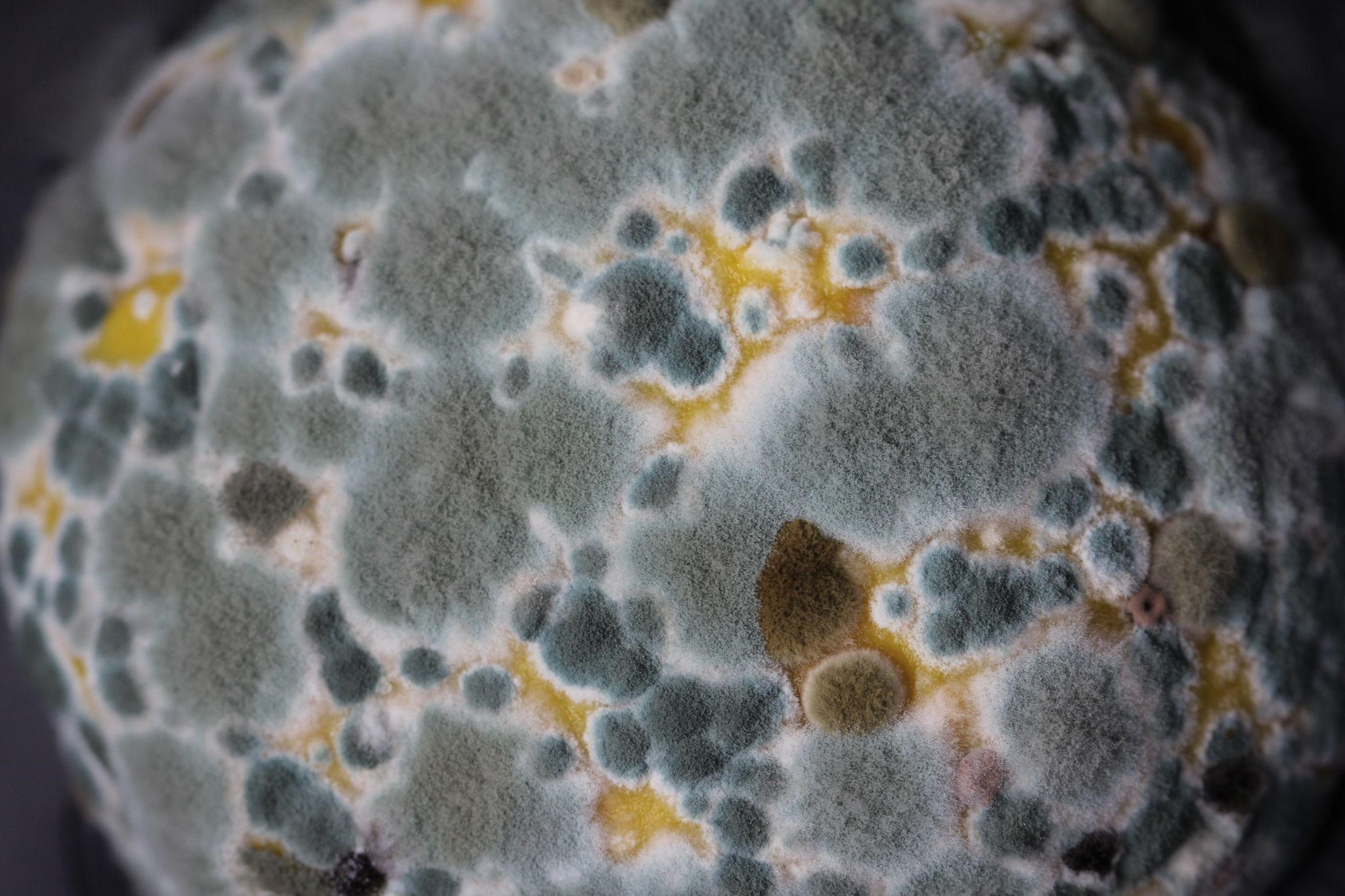
The mushroom green mold disease caused by pathogenic fungi Trichoderma spp. is the most harmful disease for mushroom production. The name “green mold” refers to the dark green color of sporulating colonies. It can cause yield losses between 60% and 100%, and thus serious economic loss. The green mold disease is especially problematic in organic mushroom production as organic farming prohibits fungicides treatment. The main problem is belated detection of the green mold disease when the damage to the yield is irreversible. EkoFungi (https://www.systemekofungi.com), a SME where our DRAGON researchers performed their residencies, is faced with this problem (described in more detail in https://datadragon.eu/2022/01/10/multidisciplinary-approach-combining-metagenomics-molecular-biology-and-hyperspectral-imaging-applied-for-detection-of-fungal-pathogens-on-mushroom-farm/). Thus, the development of an early detection strategy of green mold disease causative agent, Trichoderma spp. is an imperative and should start with an in-depth study of the diversity of compost and casing microbial communities.
Simultaneous analysis of the microbial community as a whole can be achieved using a metagenomic approach. Metagenomics involves the analysis of the collective genome of biological communities from an environmental sample. It helps to identify the species present in the community and can provide insights about the metabolic activities and functional roles of the microbes in the environmental sample. Amplicon-based metagenomic analysis produces big biodiversity data using bulk sample-based DNA metabarcoding. DNA metabarcoding of fungi uses the internal transcribed spacer region (ITS) of the nuclear ribosomal DNA to make community-wide taxonomic assignments. Using genomic DNA extracted from the compost and casing samples, as well as suitable primers for ITS region amplification, it is possible to produce sequence data necessary for subsequent taxonomic identification of present fungal communities, while multiple, time dependent sampling gives insights into the fungal community diversity changes during period of mushroom growth. In this way it is possible to monitor how microbial communities change over time and depending on developmental phase, as well as to detect a time point when pathogenic fungi such as Trichoderma start to grow.
One of the main goals of the DRAGON project is to enhance the scientific and technological capacity of BioSense researchers to perform analyses of multiscale multisource data that include metagenomic data. This is why our researchers set out to perform metagenomic analyses on real-life samples, collected during the residencies in the Serbian SME Ekofungi, a company with organic certified mushroom production in Serbia. The results of analyses were presented on one of the DRAGON workshops (described in more detail in https://datadragon.eu/2022/01/10/multidisciplinary-approach-combining-metagenomics-molecular-biology-and-hyperspectral-imaging-applied-for-detection-of-fungal-pathogens-on-mushroom-farm/ ).
Data on fungal community composition and abundance in the samples (Fig.1) are produced using bioinformatic processing of DNA metabarcoding data. Because of data quantity this kind of analysis is computationally intensive and demands at least basic bioinformatic experience. The raw sequence data should be firstly merged and filtered to obtain Clean Data ready for subsequent analysis. During the next steps of analysis Operational Taxonomic Units (OTUs) clustering is carried out based on effective data. OTUs are then used in further diversity analysis and comparisons of community compositions between samples. As a result, taxonomic composition of fungal communities in compost and casing soil from champignon production has been resolved and eight Trichoderma species among which T. harzianum and T. aggressivum as the most important causative agents of green mold disease has been identified.
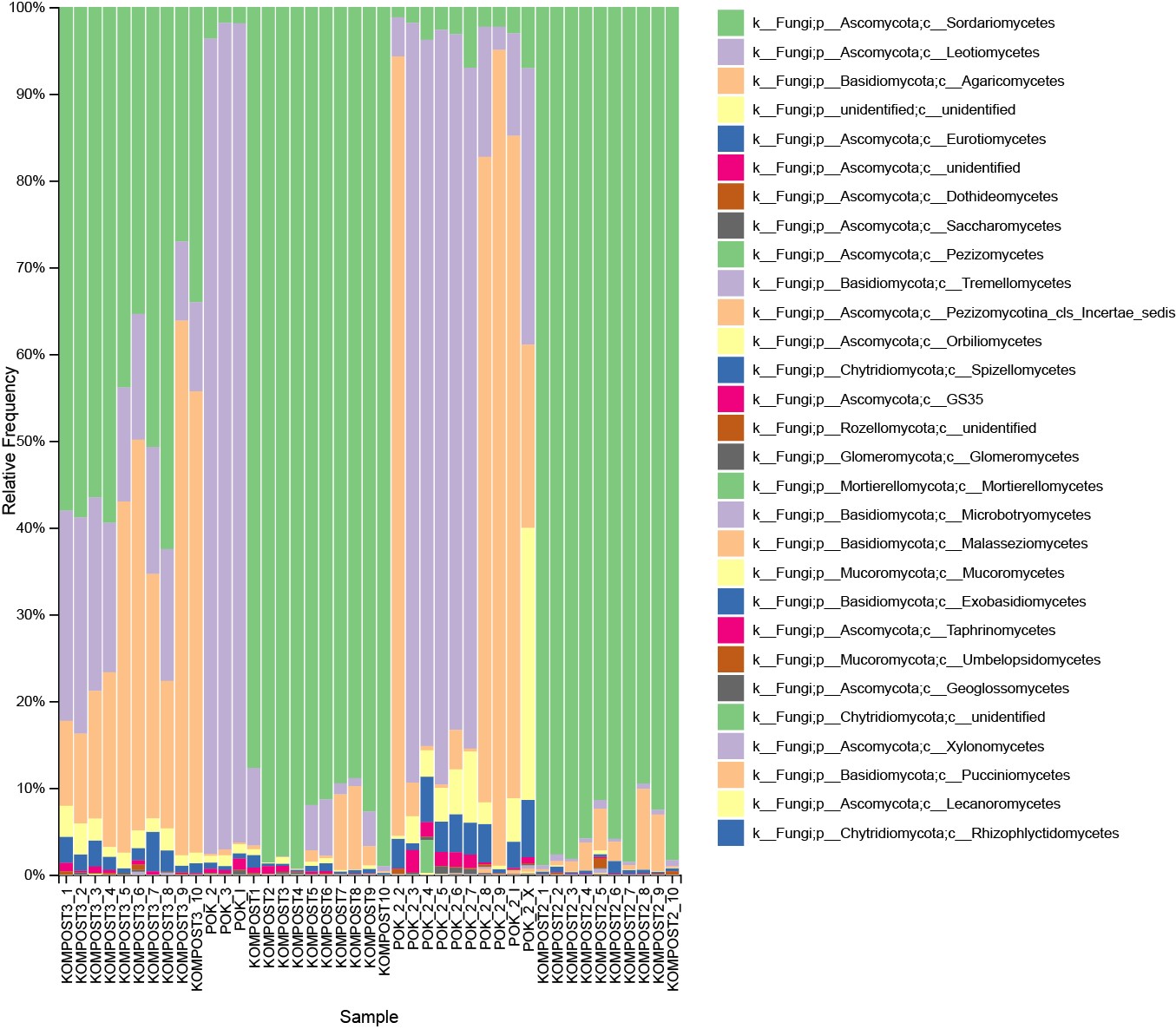
Figure 1. Taxonomic diversity bar plot for 43 compost and casing soil samples that we collected in EkoFungi company for organic mushroom production.